About the CoMSES Model Library more info
Our mission is to help computational modelers at all levels engage in the establishment and adoption of community standards and good practices for developing and sharing computational models. Model authors can freely publish their model source code in the Computational Model Library alongside narrative documentation, open science metadata, and other emerging open science norms that facilitate software citation, reproducibility, interoperability, and reuse. Model authors can also request peer review of their computational models to receive a DOI.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with additional detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 10 of 83 results for "disease%20infection%20epidemic%20epidemiology%20pandemic" clear search
Peer reviewed MOOvPOPsurveillance
Matthew Gompper Aniruddha Belsare Joshua J Millspaugh | Published Tuesday, April 04, 2017 | Last modified Tuesday, May 12, 2020MOOvPOPsurveillance was developed as a tool for wildlife agencies to guide collection and analysis of disease surveillance data that relies on non-probabilistic methods like harvest-based sampling.
The Effect of Different Governmental Pandemic Control Measures on the Spread of a Virus Disease
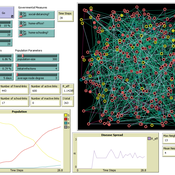
Chiara Letter | Published Wednesday, January 26, 2022In this model, the spread of a virus disease in a network consisting of school pupils, employed, and umemployed people is simulated. The special feature in this model is the distinction between different types of links: family-, friends-, school-, or work-links. In this way, different governmental measures can be implemented in order to decelerate or stop the transmission.
Hybrid Agent-Based and Equation Based Model for Infectious Disease Spread
Elizabeth Hunter Brian Mac Namee John Kelleher | Published Sunday, April 19, 2020Our model is hybrid agent-based and equation based model for human air-borne infectious diseases measles. It follows an SEIR (susceptible, exposed,infected, and recovered) type compartmental model with the agents moving be-tween the four state relating to infectiousness. However, the disease model canswitch back and forth between agent-based and equation based depending onthe number of infected agents. Our society model is specific using the datato create a realistic synthetic population for a county in Ireland. The modelincludes transportation with agents moving between their current location anddesired destination using predetermined destinations or destinations selectedusing a gravity model.
Diffusion dynamics in small-world networks with heterogeneous consumers
Sebastiano Delre | Published Saturday, September 10, 2011 | Last modified Saturday, April 27, 2013This model simulates diffusion curves and it allows to test how social influence, network structure and consumer heterogeneity affect their spreads and their speeds.
Peer reviewed MHMSLeptoDy (Multi-host, multi-serovar Leptospira Dynamics Model)
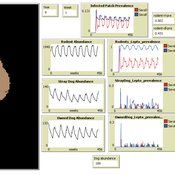
Aniruddha Belsare Matthew Gompper Meghan Mason Claudia Munoz-Zanzi | Published Tuesday, January 29, 2019 | Last modified Tuesday, March 12, 2019Leptospirosis is a neglected, bacterial zoonosis with worldwide distribution, primarily a disease of poverty. More than 200 pathogenic serovars of Leptospira bacteria exist, and a variety of species may act as reservoirs for these serovars. Human infection is the result of direct or indirect contact with Leptospira bacteria in the urine of infected animal hosts, primarily livestock, dogs, and rodents. There is increasing evidence that dogs and dog-adapted serovar Canicola play an important role in the burden of leptospirosis in humans in marginalized urban communities. What is needed is a more thorough understanding of the transmission dynamics of Leptospira in these marginalized urban communities, specifically the relative importance of dogs and rodents in the transmission of Leptospira to humans. This understanding will be vital for identifying meaningful intervention strategies.
One of the main objectives of MHMSLeptoDy is to elucidate transmission dynamics of host-adapted Leptospira strains in multi-host system. The model can also be used to evaluate alternate interventions aimed at reducing human infection risk in small-scale communities like urban slums.
Peer reviewed COMOKIT
Patrick Taillandier Alexis Drogoul Benoit Gaudou Kevin Chapuis Nghi Huyng Quang Doanh Nguyen Ngoc Arthur Brugière Pierre Larmande Marc Choisy Damien Philippon | Published Tuesday, May 26, 2020 | Last modified Wednesday, July 01, 2020In the face of the COVID-19 pandemic, public health authorities around the world have experimented, in a short period of time, with various combinations of interventions at different scales. However, as the pandemic continues to progress, there is a growing need for tools and methodologies to quickly analyze the impact of these interventions and answer concrete questions regarding their effectiveness, range and temporality.
COMOKIT, the COVID-19 modeling kit, is such a tool. It is a computer model that allows intervention strategies to be explored in silico before their possible implementation phase. It can take into account important dimensions of policy actions, such as the heterogeneity of individual responses or the spatial aspect of containment strategies.
In COMOKIT, built using the agent-based modeling and simulation platform GAMA, the profiles, activities and interactions of people, person-to-person and environmental transmissions, individual clinical statuses, public health policies and interventions are explicitly represented and they all serve as a basis for describing the dynamics of the epidemic in a detailed and realistic representation of space.
…
Peer reviewed MIOvPOPsurveillance
Aniruddha Belsare | Published Monday, April 13, 2020MIOvPOPsurveillance is set up to simulate harvest-based chronic wasting disease (CWD) surveillance of white-tailed deer (Odocoileus virginianus) populations in select Michigan Counties. New regions can be readily added, also the model can be readily adapted for other disease systems and used for informed-decision making during planning and implementation stages of disease surveillance in wildlife and free-ranging species.
A Modelling4All/NetLogo model of the Spanish Flu Pandemic
Ken Kahn | Published Monday, August 05, 2013 | Last modified Monday, August 05, 2013A global model of the 1918-19 Influenza Pandemic. It can be run to match history or explore counterfactual questions about the influence of World War I on the dynamics of the epidemic. Explores two theories of the location of the initial infection.
Potato late blight model
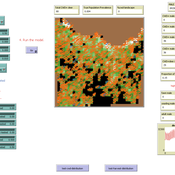
Francine Pacilly | Published Friday, April 13, 2018The purpose of the model is to simulate the spatial dynamics of potato late blight to analyse whether resistant varieties can be used effectively for sustainable disease control. The model represents an agricultural landscape with potato fields and data of a Dutch agricultural region is used as input for the model. We simulated potato production, disease spread and pathogen evolution during the growing season (April to September) for 36 years. Since late blight development and crop growth is weather dependent, measured weather data is used as model input. A susceptible and late blight resistant potato variety are distinguished. The resistant variety has a potentially lower yield but cannot get infected with the disease. However, during the growing season virulent spores can emerge as a result of mutations during spore production. This new virulent strain is able to infect the resistant fields, resulting in resistance breakdown. The model shows how disease severity, resistance durability and potato yield are affected by the fraction of fields across a landscape with a disease-resistant potato variety.
Simulating the Transmission of Foot-And-Mouth Disease Among Mobile Herds in the Far North Region, Cameroon
Mark Moritz Hyeyoung Kim Ningchuan Xiao Rebecca Garabed Laura W Pomeroy | Published Wednesday, April 13, 2016This model simulates movements of mobile pastoralists and their impacts on the transmission of foot-and-mouth disease (FMD) in the Far North Region of Cameroon.
Displaying 10 of 83 results for "disease%20infection%20epidemic%20epidemiology%20pandemic" clear search