About the CoMSES Model Library more info
Our mission is to help computational modelers at all levels engage in the establishment and adoption of community standards and good practices for developing and sharing computational models. Model authors can freely publish their model source code in the Computational Model Library alongside narrative documentation, open science metadata, and other emerging open science norms that facilitate software citation, reproducibility, interoperability, and reuse. Model authors can also request peer review of their computational models to receive a DOI.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with additional detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 10 of 11 results epidemiology clear search
A simple computational algorithm for simulating population substance use
Jacob Borodovsky | Published Thursday, March 27, 2025This code simulates individual-level, longitudinal substance use patterns that can be used to understand how cross-sectional U-shaped distributions of population substance use emerge. Each independent computational object transitions between two states: using a substance (State 1), or not using a substance (State 2). The simulation has two core components. Component 1: each object is assigned a unique risk factor transition probability and unique protective factor transition probability. Component 2: each object’s current decision to use or not use the substance is influenced by the object’s history of decisions (i.e., “path dependence”).
epiworldR Type: Fast Agent-Based Epi Models
George G. Vega Yon Derek Meyer | Published Monday, August 26, 2024A flexible framework for Agent-Based Models (ABM), the ‘epiworldR’ package provides methods for prototyping disease outbreaks and transmission models using a ‘C++’ backend, making it very fast. It supports multiple epidemiological models, including the Susceptible-Infected-Susceptible (SIS), Susceptible-Infected-Removed (SIR), Susceptible-Exposed-Infected-Removed (SEIR), and others, involving arbitrary mitigation policies and multiple-disease models. Users can specify infectiousness/susceptibility rates as a function of agents’ features, providing great complexity for the model dynamics. Furthermore, ‘epiworldR’ is ideal for simulation studies featuring large populations.
The Levers of HIV Model
Arthur Hjorth Wouter Vermeer C. Hendricks Brown Uri Wilensky Can Gurkan | Published Tuesday, March 08, 2022 | Last modified Tuesday, October 31, 2023Chicago’s demographic, neighborhood, sex risk behaviors, sexual network data, and HIV prevention and treatment cascade information from 2015 were integrated as input to a new agent-based model (ABM) called the Levers-of-HIV-Model (LHM). This LHM, written in NetLogo, forms patterns of sexual relations among Men who have Sex with Men (MSM) based on static traits (race/ethnicity, and age) and dynamic states (sexual relations and practices) that are found in Chicago. LHM’s five modules simulate and count new infections at the two marker years of 2023 and 2030 for a wide range of distinct scenarios or levers, in which the levels of PrEP and ART linkage to care, retention, and adherence or viral load are increased over time from the 2015 baseline levels.
Peer reviewed Virus Transmission with Super-spreaders
J M Applegate | Published Saturday, September 11, 2021A curious aspect of the Covid-19 pandemic is the clustering of outbreaks. Evidence suggests that 80\% of people who contract the virus are infected by only 19% of infected individuals, and that the majority of infected individuals faile to infect another person. Thus, the dispersion of a contagion, $k$, may be of more use in understanding the spread of Covid-19 than the reproduction number, R0.
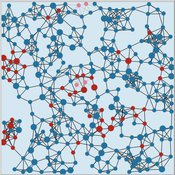
The Virus Transmission with Super-spreaders model, written in NetLogo, is an adaptation of the canonical Virus Transmission on a Network model and allows the exploration of various mitigation protocols such as testing and quarantines with both homogenous transmission and heterogenous transmission.
The model consists of a population of individuals arranged in a network, where both population and network degree are tunable. At the start of the simulation, a subset of the population is initially infected. As the model runs, infected individuals will infect neighboring susceptible individuals according to either homogenous or heterogenous transmission, where heterogenous transmission models super-spreaders. In this case, k is described as the percentage of super-spreaders in the population and the differing transmission rates for super-spreaders and non super-spreaders. Infected individuals either recover, at which point they become resistant to infection, or die. Testing regimes cause discovered infected individuals to quarantine for a period of time.
Holmestrand School Model
Jessica Dimka | Published Friday, June 18, 2021 | Last modified Friday, April 29, 2022The Holmestrand model is an epidemiological agent-based model. Its aim is to test hypotheses related to how the social and physical environment of a residential school for children with disabilities might influence the spread of an infectious disease epidemic among students and staff. Annual reports for the Holmestrand School for the Deaf (Norway) are the primary sources of inspiration for the modeled school, with additional insights drawn from other archival records for schools for children with disabilities in early 20th century Norway and data sources for the 1918 influenza pandemic. The model environment consists of a simplified boarding school that includes residential spaces for students and staff, classrooms, a dining room, common room, and an outdoor area. Students and staff engage in activities reflecting hourly schedules suggested by school reports. By default, a random staff member is selected as the first case and is infected with disease. Subsequent transmission is determined by agent movement and interactions between susceptible and infectious pairs.
AIforGoodSimulator - Modeling Covid-19 Spread and Potential Interventions in Refugee Camps
Shyaam Ramkumar Woi Sok Oh | Published Thursday, March 18, 2021The Netlogo model is a conceptualization of the Moria refugee camp, capturing the household demographics of refugees in the camp, a theoretical friendship network based on values, and an abstraction of their daily activities. The model then simulates how Covid-19 could spread through the camp if one refugee is exposed to the virus, utilizing transmission probabilities and the stages of disease progression of Covid-19 from susceptible to exposed to asymptomatic / symptomatic to mild / severe to recovered from literature. The model also incorporates various interventions - PPE, lockdown, isolation of symptomatic refugees - to analyze how they could mitigate the spread of the virus through the camp.
Classical Swine Fever in wild boars
Volker Grimm Stephanie Kramer-Schadt Cédric Scherer Martin Lange Hans-Hermann Thulke | Published Friday, September 06, 2019The model is a combination of a spatially explicit, stochastic, agent-based model for wild boars (Sus scrofa L.) and an epidemiological model for the Classical Swine Fever (CSF) virus infecting the wild boars.
The original model (Kramer-Schadt et al. 2009) was used to assess intrinsic (system immanent host-pathogen interaction and host life-history) and extrinsic (spatial extent and density) factors contributing to the long-term persistence of the disease and has further been used to assess the effects of intrinsic dynamics (Lange et al. 2012a) and indirect transmission (Lange et al. 2016) on the disease course. In an applied context, the model was used to test the efficiency of spatiotemporal vaccination regimes (Lange et al. 2012b) as well as the risk of disease spread in the country of Denmark (Alban et al. 2005).
References: See ODD model description.
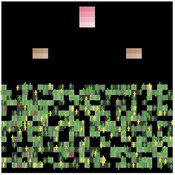
Mission San Diego Model
Carolyn Orbann | Published Monday, April 15, 2019The Mission San Diego model is an epidemiological model designed to test hypotheses related to the spread of the 1805-1806 measles epidemic among indigenous residents of Mission San Diego during the early mission period in Alta California. The model community is based on the population of the Mission San Diego community, as listed in the parish documents (baptismal, marriage, and death records). Model agents are placed on a map-like grid that consists of houses, the mission church, a women’s dormitory (monjeria) adjacent to the church, a communal kitchen, priest’s quarters, and agricultural fields. They engage in daily activities that reflect known ethnographic patterns of behavior at the mission. A pathogen is introduced into the community and then it spreads throughout the population as a consequence of individual agent movements and interactions.
St Anthony flu
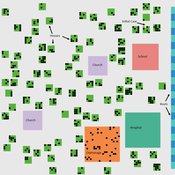
Lisa Sattenspiel | Published Monday, April 15, 2019The St Anthony flu model is an epidemiological model designed to test hypotheses related to the spread of the 1918 influenza pandemic among residents of a small fishing community in Newfoundland and Labrador. The 1921 census data from Newfoundland and Labrador are used to ensure a realistic model population; the community of St. Anthony, NL, located on the tip of the Northern Peninsula of the island of Newfoundland is the specific population modeled. Model agents are placed on a map-like grid that consists of houses, two churches, a school, an orphanage, a hospital, and several boats. They engage in daily activities that reflect known ethnographic patterns of behavior in St. Anthony and other similar communities. A pathogen is introduced into the community and then it spreads throughout the population as a consequence of individual agent movements and interactions.
CA-MRSA Demonstration Model
Jonathan Ozik Charles Macal Kenneth Letendre Irene Lee | Published Tuesday, January 06, 2015We demonstrate how a simple model of community associated Methicillin-resistant Staphylococcus aureus (CA-MRSA) can be easily constructed by leveraging the statecharts and ReLogo capabilities in Repast Simphony.
Displaying 10 of 11 results epidemiology clear search