About the CoMSES Model Library more info
Our mission is to help computational modelers at all levels engage in the establishment and adoption of community standards and good practices for developing and sharing computational models. Model authors can freely publish their model source code in the Computational Model Library alongside narrative documentation, open science metadata, and other emerging open science norms that facilitate software citation, reproducibility, interoperability, and reuse. Model authors can also request peer review of their computational models to receive a DOI.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with additional detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 10 of 114 results for "disease infection epidemic epidemiology pandemic" clear search
Model of Rental Evictions in Phoenix During the Covid-19 Pandemic
Sean Bergin J M Applegate | Published Saturday, July 31, 2021 | Last modified Friday, October 15, 2021The purpose of this model is to explore the dynamics of residency and eviction for households renting in the greater Phoenix (Arizona) metropolitan area. The model uses a representative population of renters modified from American Community Survey (ACS) data that includes demographic, housing and economic information. Each month, households pay their subsistence, rental and utility bills. If a household is unable to pay their monthly rent or utility bill they apply for financial assistance. This model provides a platform to understand the impact of various economic shock upon households. Also, the model includes conditions that occurred as a result of the Covid-19 pandemic which allows for the study of eviction mitigation strategies that were employed, such as the eviction moratorium and stimulus payments. The model allows us to make preliminary predictions concerning the number of households that may be evicted once the moratorium on evictions ends and the long-term effects on the number of evicted households in the greater Phoenix area going forward.
Peer reviewed INOvCWD
Aniruddha Belsare | Published Wednesday, June 01, 2022 | Last modified Wednesday, July 10, 2024INOvCWD is a spatially-explicit, agent-based model designed to simulate the spread of chronic wasting disease (CWD) in Indiana’s white-tailed deer populations.
Evolution of Conditional Cooperation
M Manning Marco Janssen Oyita Udiani | Published Thursday, August 01, 2013 | Last modified Friday, May 13, 2022Cultural group selection model used to evaluate the conditions for agents to evolve who have other-regarding preferences in making decisions in public good games.
Pastoralscape
Matthew Sottile | Published Tuesday, October 12, 2021Pastoralscape is a model of human agents, lifestock health and contageous disease for studying the impact of human decision making in pastoral communities within East Africa on livestock populations. It implements an event-driven agent based model in Python 3.
Food Safety Inspection Model - Stores Signal with Errors
Sara Mcphee-Knowles | Published Wednesday, March 05, 2014 | Last modified Monday, April 08, 2019The Inspection Model represents a basic food safety system where inspectors, consumers and stores interact. The purpose of the model is to provide insight into an optimal level of inspectors in a food system by comparing three search strategies.
A consumer-demand simulation for Smart Metering tariffs (Innovation Diffusion)
Martin Rixin | Published Thursday, August 18, 2011 | Last modified Saturday, April 27, 2013An Agent-based model simulates consumer demand for Smart Metering tariffs. It utilizes the Bass Diffusion Model and Rogers´s adopter categories. Integration of empirical census microdata enables a validated socio-economic background for each consumer.
An epidemiological-economic agent-based model of Huanglongbing and Asian citrus psyllid control in California
Jonathan Kaplan | Published Wednesday, June 01, 2022This model simulates economic and epidemiological interaction between citrus production and the disease Huanglongbing (HLB), which is vectored by the Asian citrus psyllid. The model is used to evaluate area-wide coordinated spraying when free-riding is possible given individuals’ beliefs in other grower participation in area-wide spraying and in the information provided by extension on the threat as HLB spread.
Mission San Diego Model
Carolyn Orbann | Published Monday, April 15, 2019The Mission San Diego model is an epidemiological model designed to test hypotheses related to the spread of the 1805-1806 measles epidemic among indigenous residents of Mission San Diego during the early mission period in Alta California. The model community is based on the population of the Mission San Diego community, as listed in the parish documents (baptismal, marriage, and death records). Model agents are placed on a map-like grid that consists of houses, the mission church, a women’s dormitory (monjeria) adjacent to the church, a communal kitchen, priest’s quarters, and agricultural fields. They engage in daily activities that reflect known ethnographic patterns of behavior at the mission. A pathogen is introduced into the community and then it spreads throughout the population as a consequence of individual agent movements and interactions.
Covid-19-Belief-network-Hybrid-Model
Morteza Mahmoudzadeh | Published Sunday, September 05, 2021Digital social networks facilitate the opinion dynamics and idea flow and also provide reliable data to understand these dynamics. Public opinion and cooperation behavior are the key factors to determine the capacity of a successful and effective public policy. In particular, during the crises, such as the Corona virus pandemic, it is necessary to understand the people’s opinion toward a policy and the performance of the governance institutions. The problem of the mathematical explanation of the human behaviors is to simplify and bypass some of the essential process. To tackle this problem, we adopted a data-driven strategy to extract opinion and behavioral patterns from social media content to reflect the dynamics of society’s average beliefs toward different topics. We extracted important subtopics from social media contents and analyze the sentiments of users at each subtopic. Subsequently, we structured a Bayesian belief network to demonstrate the macro patters of the beliefs, opinions, information and emotions which trigger the response toward a prospective policy. We aim to understand the factors and latent factors which influence the opinion formation in the society. Our goal is to enhance the reality of the simulations. To capture the dynamics of opinions at an artificial society we apply agent-based opinion dynamics modeling. We intended to investigate practical implementation scenarios of this framework for policy analysis during Corona Virus Pandemic Crisis. The implemented modular modeling approach could be used as a flexible data-driven policy making tools to investigate public opinion in social media. The core idea is to put the opinion dynamics in the wider contexts of the collective decision-making, data-driven policy-modeling and digital democracy. We intended to use data-driven agent-based modeling as a comprehensive analysis tools to understand the collective opinion dynamics and decision making process on the social networks and uses this knowledge to utilize network-enabled policy modeling and collective intelligence platforms.
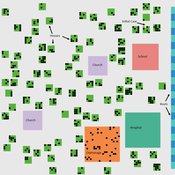
St Anthony flu
Lisa Sattenspiel | Published Monday, April 15, 2019The St Anthony flu model is an epidemiological model designed to test hypotheses related to the spread of the 1918 influenza pandemic among residents of a small fishing community in Newfoundland and Labrador. The 1921 census data from Newfoundland and Labrador are used to ensure a realistic model population; the community of St. Anthony, NL, located on the tip of the Northern Peninsula of the island of Newfoundland is the specific population modeled. Model agents are placed on a map-like grid that consists of houses, two churches, a school, an orphanage, a hospital, and several boats. They engage in daily activities that reflect known ethnographic patterns of behavior in St. Anthony and other similar communities. A pathogen is introduced into the community and then it spreads throughout the population as a consequence of individual agent movements and interactions.
Displaying 10 of 114 results for "disease infection epidemic epidemiology pandemic" clear search