About the CoMSES Model Library more info
Our mission is to help computational modelers at all levels engage in the establishment and adoption of community standards and good practices for developing and sharing computational models. Model authors can freely publish their model source code in the Computational Model Library alongside narrative documentation, open science metadata, and other emerging open science norms that facilitate software citation, reproducibility, interoperability, and reuse. Model authors can also request peer review of their computational models to receive a DOI.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with additional detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 6 of 6 results for 'Stephanie Dornschneider'
Animal territory formation (Reusable Building Block RBB)
Robert Zakrzewski Stephanie Kramer-Schadt Volker Grimm | Published Sunday, November 12, 2023This is a generic sub-model of animal territory formation. It is meant to be a reusable building block, but not in the plug-and-play sense, as amendments are likely to be needed depending on the species and region. The sub-model comprises a grid of cells, reprenting the landscape. Each cell has a “quality” value, which quantifies the amount of resources provided for a territory owner, for example a tiger. “Quality” could be prey density, shelter, or just space. Animals are located randomly in the landscape and add grid cells to their intial cell until the sum of the quality of all their cells meets their needs. If a potential new cell to be added is owned by another animal, competition takes place. The quality values are static, and the model does not include demography, i.e. mortality, mating, reproduction. Also, movement within a territory is not represented.
Survival analysis and Population viability analysis of the Northern Bald Ibis
Sinah Drenske Viktoriia Radchuk Cédric Scherer Corinna Esterer Ingo Kowarik Johannes Fritz Stephanie Kramer-Schadt | Published Thursday, August 04, 2022This is the full repository to run the survival analysis (in R) and run the population viability model and its analysis (NetLogo + R) of the Northern Bald Ibis (NBI) presented in the study
On the road to self-sustainability: Reintroduced migratory European Northern Bald Ibises (Geronticus eremita) still need management interventions for population viability
by Sinah Drenske, Viktoriia Radchuk, Cédric Scherer, Corinna Esterer, Ingo Kowarik, Johannes Fritz, Stephanie Kramer-Schadt
…
Modeling the emergence of migratory corridors and foraging hotspots of the green sea turtle
Mayeul Dalleau Volker Grimm Jérôme Bourjea Stephanie Kramer-Schadt | Published Friday, April 05, 2019 | Last modified Tuesday, September 17, 2019The model represents migration of the green sea turtle, Chelonia mydas, between foraging and breeding sites in the Southwest Indian Ocean. The purpose of the model is to investigate the impact of local environmental conditions, including the quality of foraging sites and ocean currents, on emerging migratory corridors and reproductive output and to thereby identify conservation priority sites.
Corresponding article to found here: https://onlinelibrary.wiley.com/doi/epdf/10.1002/ece3.5552
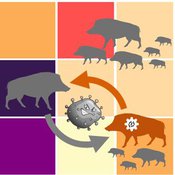
Classical Swine Fever in wild boars
Cédric Scherer Martin Lange Volker Grimm Hans-Hermann Thulke Stephanie Kramer-Schadt | Published Friday, September 06, 2019The model is a combination of a spatially explicit, stochastic, agent-based model for wild boars (Sus scrofa L.) and an epidemiological model for the Classical Swine Fever (CSF) virus infecting the wild boars.
The original model (Kramer-Schadt et al. 2009) was used to assess intrinsic (system immanent host-pathogen interaction and host life-history) and extrinsic (spatial extent and density) factors contributing to the long-term persistence of the disease and has further been used to assess the effects of intrinsic dynamics (Lange et al. 2012a) and indirect transmission (Lange et al. 2016) on the disease course. In an applied context, the model was used to test the efficiency of spatiotemporal vaccination regimes (Lange et al. 2012b) as well as the risk of disease spread in the country of Denmark (Alban et al. 2005).
References: See ODD model description.
A Simulation of Arab Spring Protests Informed by Qualitative Evidence
Bruce Edmonds Stephanie Dornschneider | Published Monday, April 29, 2019 | Last modified Friday, May 24, 2019The purpose of the simulation was to explore and better understand the process of bridging between an analysis of qualitative data and the specification of a simulation. This may be developed for more serious processes later but at the moment it is merely an illustration.
This exercise was done by Stephanie Dornschneider (School of Politics and International Relations, University College Dublin) and Bruce Edmonds to inform the discussion at the Lorentz workshop on “Integrating Qualitative and Quantitative Data using Social Simulation” at Leiden in April 2019. The qualitative data was collected and analysed by SD. The model specification was developed as the result of discussion by BE & SD. The model was programmed by BE. This is described in a paper submitted to Social Simulation 2019 and (to some extent) in the slides presented at the workshop.
Value Chain Marketing (VCM)
Stephanie Hintze | Published Monday, April 14, 2014 | Last modified Thursday, October 16, 2014Inspired by the SKIN model, the basic concept here is to model the acceptance and implementation of supplier innovations. This model includes three types of agents comprising suppliers, manufacturers and applicators.