About the CoMSES Model Library more info
Our mission is to help computational modelers at all levels engage in the establishment and adoption of community standards and good practices for developing and sharing computational models. Model authors can freely publish their model source code in the Computational Model Library alongside narrative documentation, open science metadata, and other emerging open science norms that facilitate software citation, reproducibility, interoperability, and reuse. Model authors can also request peer review of their computational models to receive a DOI.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with additional detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 10 of 109 results for "Marc Deconchat" clear search
MoPAgrIB: simulating savannah landscape mosaic under shifting cultivation
Nicolas Becu Marc Deconchat Eric Garine Kouami Kokou Christine Raimond | Published Monday, May 27, 2013 | Last modified Tuesday, January 21, 2014MoPAgrIB model simulates the movement of cultivated patches in a savannah vegetation mosaic ; how they move and relocate through the landscape, depending on farming practices, population growth, social rules and vegetation growth.
Port of Mars simplified
Marco Janssen | Published Tuesday, January 14, 2020This is a simulation model to explore possible outcomes of the Port of Mars cardgame. Port of Mars is a resource allocation game examining how people navigate conflicts between individual goals and common interests relative to shared resources. The game involves five players, each of whom must decide how much of their time and effort to invest in maintaining public infrastructure and renewing shared resources and how much to expend in pursuit of their individual goals. In the game, “Upkeep” is a number that represents the physical health of the community. This number begins at 100 and goes down by twenty-five points each round, representing resource consumption and wear and tear on infrastructure. If that number reaches zero, the community collapses and everyone dies.
Population dynamics
Marco Janssen | Published Monday, December 31, 2018Simple population dynamics model used in Introduction to Agent-Based Modeling by Marco Janssen. For more information see https://intro2abm.com/
Diffusion of innovations
Marco Janssen | Published Tuesday, January 14, 20203 simple models to illustrate diffusion of innovations.
The models are discussed in Introduction to Agent-Based Modeling by Marco Janssen. For more information see https://intro2abm.com/
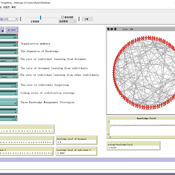
How to Manage Individual Forgetting
wiseyanjie | Published Wednesday, July 17, 2019we extend the basic simulation model of March by incorporating forgetting and three knowledge management strategies—personalization, codification, and mixed—to explore the impacts of different knowledge management strategies and forgetting on organizational knowledge level.
Governing the commons
Marco Janssen | Published Tuesday, January 14, 2020 | Last modified Sunday, July 17, 2022Model on the use of shared renewable resources including impact of imitation via success-bias and altruistic punishment.
The model is discussed in Introduction to Agent-Based Modeling by Marco Janssen. For more information see https://intro2abm.com/
Peer reviewed Artificial Anasazi
Marco Janssen | Published Tuesday, September 07, 2010 | Last modified Saturday, April 27, 2013Replication of the well known Artificial Anasazi model that simulates the population dynamics between 800 and 1350 in the Long House Valley in Arizona.
Consumats on a network
Marco Janssen | Published Tuesday, January 14, 2020 | Last modified Tuesday, May 30, 2023Consumer agents make choices which products to choose using the consumat approach. In this approach agents will make choices using deliberation, repetition, imitation or social comparison dependent on the level of need satisfaction and uncertainty.
The model is discussed in Introduction to Agent-Based Modeling by Marco Janssen. For more information see https://intro2abm.com/
Peer reviewed The Garbage Can Model of Organizational Choice
Guido Fioretti | Published Monday, April 20, 2020 | Last modified Thursday, April 23, 2020The Garbage Can Model of Organizational Choice is a fundamental model of organizational decision-making originally proposed by J.D. Cohen, J.G. March and J.P. Olsen in 1972. In the 2000s, G. Fioretti and A. Lomi presented a NetLogo agent-based interpretation of this model. This code is the NetLogo 6.1.1 updated version of the Fioretti-Lomi model.
Rangeland and evolution of management styles
Marco Janssen | Published Tuesday, January 14, 2020Provided is a landscape of properties where pastoralists make decisions how much livestock they put on their property and how much to suppress fire from occuring. Rangelands can be grass dominated, or unproductive shrubb dominated. Overgrazing and fire suppresion lead to shrub dominated landscapes. What management strategies evolve, and how is this impacted by policies?
The model is discussed in Introduction to Agent-Based Modeling by Marco Janssen. For more information see https://intro2abm.com/.
Displaying 10 of 109 results for "Marc Deconchat" clear search