About the CoMSES Model Library more info
Our mission is to help computational modelers at all levels engage in the establishment and adoption of community standards and good practices for developing and sharing computational models. Model authors can freely publish their model source code in the Computational Model Library alongside narrative documentation, open science metadata, and other emerging open science norms that facilitate software citation, reproducibility, interoperability, and reuse. Model authors can also request peer review of their computational models to receive a DOI.
All users of models published in the library must cite model authors when they use and benefit from their code.
Please check out our model publishing tutorial and contact us if you have any questions or concerns about publishing your model(s) in the Computational Model Library.
We also maintain a curated database of over 7500 publications of agent-based and individual based models with additional detailed metadata on availability of code and bibliometric information on the landscape of ABM/IBM publications that we welcome you to explore.
Displaying 2 of 2 results military history clear search
ThomondSim
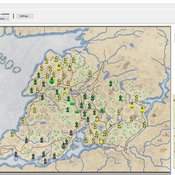
Vinicius Marino Carvalho | Published Monday, April 25, 2022 | Last modified Friday, May 12, 2023ThomondSim is a simulation of the political and economic landscape of the medieval kingdom of Thomond, southwestern Ireland, between 1276 and 1318.
Its goal is to analyze how deteriorating environmental and economic conditions caused by the Little Ice Age (LIA), the Great European Famine of 1315-1322, and wars between England and Scotland affected the outcomes of a local war involving Gaelic and English aristocratic lineages.
This ABM attempts to model both the effects of devastation on the human environment and the modus operandi of late-medieval war and diplomacy.
The model is the digital counterpart of the science discovery board game The Triumphs of Turlough. Its procedures closely correspond to the game’s mechanics, to the point that ToT can be considered an interactive, analog version of this ABM.
Gunpowder battle tactics
Xavier Rubio-Campillo Jose María Cela Francesc Xavier Hernàndez | Published Wednesday, November 20, 2013 | Last modified Tuesday, November 26, 2013This model simulates the dynamics of eighteenth-century infantry battle tactics. The goal is to explore the effect of different tactics and individual traits in the dynamics of the combat.